
PRE-ANALYTICAL PRACTICE IN MOLECULAR ASSAY IN CLINICAL AND RESEARCH SETTINGS
By: Yan Mardian
Pre-analysis refers to all the complex steps that must take place before a sample can be analyzed. Over the years, a series of studies identified that 32% to 75% of all testing errors occur in the preanalytical phase, and technological advances and quality assurance procedures have significantly reduced the number of analytic-based errors. This has exposed the pre-analysis stage as a major source of residual “error” and/or variables that can affect test results. Preanalytic factors include patient-related variables (diet, age, sex, etc.), specimen collection and labeling techniques, specimen preservatives and anticoagulants, specimen transport, and processing and storage. Potential sources of error or failure in this process include improperly ordered tests, sample misidentification, improper timing, improper fasting, improper anticoagulant/blood ratio, improper mixing, incorrect order of draw, and hemolyzed or lipemic specimens.
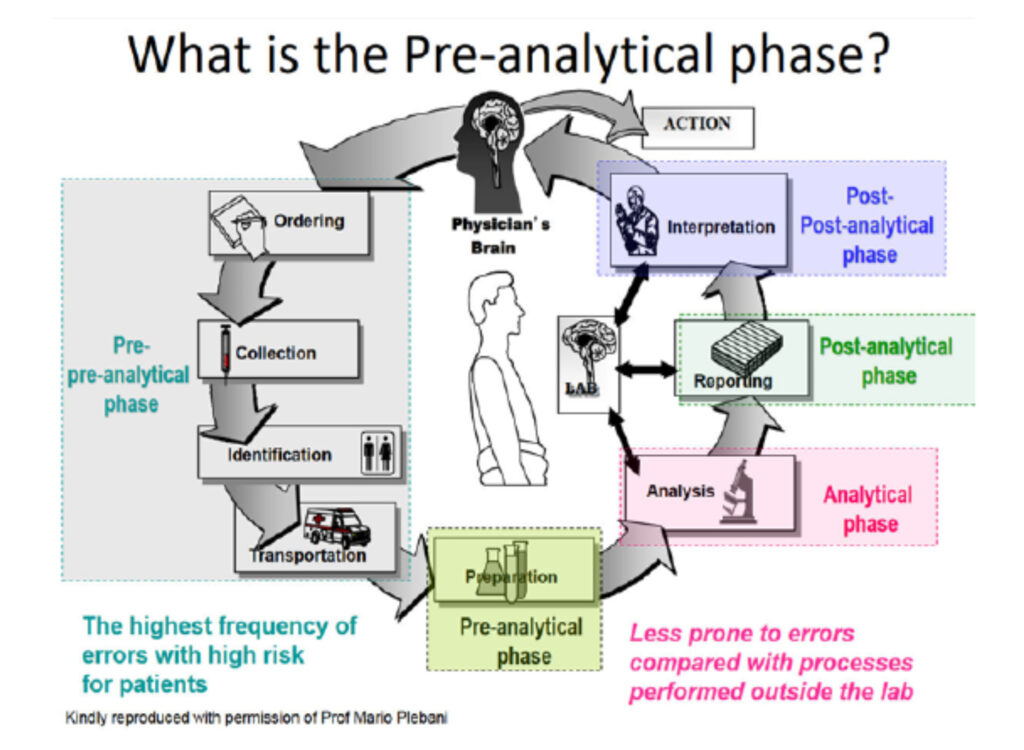
The most frequent preanalytic errors include improperly filling the sample tube, placing specimens in the wrong containers or preservatives, and selecting the incorrect test.
There is now incontrovertible evidence that the pre-analytical phase is the major source of errors in laboratory testing when used for either diagnostic or research purposes. The molecular assay is not an exception, whereby many potential preanalytical errors are similar to those occurring in other different diagnostic areas, whilst others can be classified as specific. Molecular assays for the detection of nucleic acids in biologic specimens are valuable diagnostic tools supporting clinical diagnoses and therapeutic decisions. Pre-analytical errors, contribute a significant role in common errors that take place in molecular laboratories. Certain practices in specimen collection, transportation, and storage can affect the integrity of nucleic acids before analysis. Although diagnostic errors can occur almost always and everywhere in healthcare, the vulnerability of error in molecular assays pre-analytic phase is enormously magnified when the staff is forced to work in high-throughput settings, as seen in the COVID-19 pandemic. In the case of infectious outbreaks, the generation of false-positive or false-negative test results not only jeopardizes the health of the individual patient but may also derange and disrupt the efficacy of public health policies, emergency plans, and restrictive measures established by national and international authorities for containing the outbreak.
This article addressed correct practices in specimen collection, transportation, preservation, and storage in most biological samples which are routinely received and processed in translational research laboratories. Laboratory manipulations of nucleic acids are susceptible to interferences at various stages, including specimen collection and processing. Blood and its components are among the most common samples received in molecular laboratories. Most of the received samples contain anticoagulants, therefore awareness about the possible confounding effects of each anticoagulant and selection of the proper one has paramount importance in ensuring the accuracy of reports. The introduction of inhibitory substances and contamination with false-positive signals are among the significant interferences. Currently, K2/K3-EDTA and acid citrate dextrose (ACD) are the recommended anticoagulants for molecular assays. Blood specimens for nucleic acid testing are generally collected into EDTA anticoagulant to inhibit enzymes that might break them down. Heparin is a poor choice for anticoagulant in this application because it can be coextracted with DNA and inhibits DNA polymerase in polymerase chain reactions (PCRs). There are some reports implying that heparin, even at low concentrations, may suppress DNA amplification. Overall, it is recommended to avoid the usage of a heparinized sample as much as possible in molecular assays, but in certain situations such as only one sample or emergency states, a heparinized sample should not be rejected.
Hemin from hemolysis in plasma or serum can also inhibit DNA polymerase. RNA is labile in blood or tissues; thus, these specimens must be stored appropriately by rapid freezing in liquid nitrogen if the extraction will be delayed. Extraction of nucleic acids from clinical specimens such as plasma (e.g., for viral load measurement), blood cells (e.g., for genetic testing), or tissues (e.g., for analyzing mutations in tumors) entails lysing cells and separating nucleic acids from proteins and lipids. Reagents for extraction include salts, proteases, and phenol-chloroform to denature the substances complexed with nucleic acids. This process must be optimized for specimen type to recover high-quality nucleic acids with good quantitative yield. Care must be taken to avoid contamination of specimens with target nucleic acids from other specimens or with amplified targets from specimens that have been analyzed previously in that vicinity. Accordingly, laboratories practicing nucleic acid amplification, especially PCR, should have separate preamplification, amplification, and post-amplification
The effect of freeze–thaw cycles on nucleic acid quantity and quality is under debate. As a general rule, freezing and thawing have detrimental effects on the concentration of all analytes. Ross et al. observed more than 25% decrease in blood DNA con-centration after a single freeze–thaw cycle but the quality of DNA did not change even after multiple freeze–thaw cycles. According to Chan et al., a single freeze-thaw cycle of plasma has no significant effect on DNA integrity but repeated (3 times) freezing and thawing of plasma (but not extracted DNA) leads to fragmentation of DNA. Kopreski et al. reported that a single freeze–thaw cycle results in a marked decrease of c-abl and tyrosinase mRNAs in serum (87), however a similar study by Tsui et al. showed no significant difference for RNA concentrations between untreated serum/plasma and frozen/thawed samples. Detection of nucleic acids of infectious agents by molecular methods may be individually considered. According to CLSI, for RNA viruses like human immunodeficiency virus (HIV) (and hepatitis C virus (HCV)), the plasma should be separated from whole blood into a second tube within four hours of specimen collection and WHO/UNAIDS guidelines recommend storing serum and plasma at 4°C –8°C for up to a maximum of one week. For longer storage, the specimen should be frozen at –20°C or lower. HIV is a biologically stable virus. It has been stated that HIV-1 RNA is stable in EDTA-whole blood for up to 72 hours and if the plasma is separated by centrifugation within 12 hours the genomic content of the virus can be evaluated for up to 7 days at RT.
Nasopharyngeal specimens for Respiratory viruses diagnosis (Influenza, Respiratory Syncytial Virus, Parainfluenza virus, Human Metapneumovirus, Rhinovirus, Enterovirus, Adenovirus, Coronavirus) are collected by sterile dacron/nylon swab. Nylon-flocked swabs are more efficient in comparison with other synthetic ones like rayon swabs. The most common acceptable specimen for detecting Influenza Viruses is nasopharyngeal swabs along with washes or aspirates. Throat swabs and/or nasal swabs (two sterile dry polyester swabs with aluminum or plastic shafts) are also acceptable. However, more viral loads can be obtained by nasopharyngeal swabs in comparison with oropharyngeal swabs. The swab should be transported into sterile VTM at 4°C. If the shipment is delayed by 3 to 4 days, the samples should be kept at -70°C. Sputum specimens should be obtained in a sterile container and then transported at RT for DNA analysis. If transportation time is anticipated to be longer than 30 minutes, samples should be transported to the laboratory at 4 to 8°C. The transport media that have been developed for culture may not be suitable for molecular tests and the media developed for molecular tests are not suitable for culture. The sputum specimens which are not going to be tested immediately should be refrigerated for up to 7 days. If longer storage time is needed, the specimen can be stored for a few years at -70°C. For M. tuberculosis, it is recommended to first apply N-acetyl-L-cysteine-NaOH for decontamination, then directly put thick particles of sputum on FTA cards with a foam-tipped applicator and spread over an area of 2.5 cm. Subsequently, the samples should be air-dried for 1 hour and finally put in a storage packet and stored at RT. Ethanol fixation of sputum samples does not affect the rate of M. tuberculosis detection, so the cytology slides of sputum sediments can be used for PCR detection. Stomach acidic fluid should be neutralized with phosphate buffer before DNA isolation. After the final centrifugation, the pellet can also be used for DNA isolation. . After final centrifugation, the pellet can also be used for DNA isolation.
Dried blood spots (DBSs) can be used to detect viral genomes and can be prepared on filter paper with optimal blood volumes as low as 50 μL. Even six hours lag time between blood collection and its application on DBS can lead to reliable results. The drying procedure takes 3 hours at RT in a safety cabinet (even longer in high humidity) or 1 hour at 37°C in an oven. One of the great advantages of DBS is easy handling and shipping even in rural areas with unavailable cold chain. RNA in DBS samples is stable for at least 3 months at RT. However, some studies show that RNAs are usable for as long as 1 year at RT. High

temperature (37°C) results in progressive degradation of RNA but has less effect on DNA. RNA progressive loss at 37°C is remarkable after two months. Refrigerated (in a zip bag with a desiccant pack) and frozen DBS samples can be stored for 1 year and a period of 4 years, respectively, with reliable recovery of HIV1 RNA.
Stool sample preservation in cold conditions (4°C) is recommended during transportation. Allowable transportation time at 4°C is 24-48 hours. This time is 4 hours at RT without preservation. Fixatives/preservatives such as TotalFix, Unifix, Zinc- or copper-based polyvinyl alcohol (PVA), and Ecofix can pre-serve specimens for storing and transporting at RT. DNA stabilizer can be utilized for sample preservation to protect DNA and/o RNA and prevent degradation after collection. One of the most famous stabilizers is RNAlater, which prevents RNA as well as DNA degradation at ambient temperature for one week. Another common and economic stabilizer for fecal DNA is 95% Ethanol. The procedure contains mixing 2.5 mL of 95% ethanol with approximately 1–2 g of stool. For microbiome studies, it is ideal to transport fecal specimens immediately at -20°C or -80°C. These specimens can be stored at -80°C for up to 2 years with unremarkable alteration in stool microbial composition. The storage time is reduced to a few weeks at -20°C. Viral infections can be detected in urine using PCR, using sterile containers for urine sample collection and transport at 4°C. Urine can be stored at 4°C for Epstein–Barr virus (EBV) and CMV until processing is done in one week. The specimens should be shipped on wet ice within one week for CMV evaluation. Collection of the first 10-20 mL of voided urine for HSV (first catch) and transportation within 3 hours has been recommended. The specimens can be kept at -70°C for up to 3 months, but individual manufacturer’s instructions should also be considered for storage conditions.
Reference:
- Mark S. Lifshitz. Preanalysis. in: Henry’s clinical diag-nosis and management by laboratory methods E-book. Elsevier Health Sciences; 2022. p22-34
- Lippi G, Simundic AM, Plebani M. Potential preana-lytical and analytical vulnerabilities in the laboratory diagnosis of coronavirus disease 2019 (COVID-19). Clinical Chemistry and Laboratory Medicine (CCLM). 2020 Jul 1;58(7):1070-6.
- Anvari MS, Gharib A, Abolhasani M, Azari-Yam A, Gharalari FH, Safavi M, Mirzaie AZ, Vasei M. Pre-analytical practices in the molecular diagnostic tests, a concise review. Iranian Journal of Pathology. 2021;16(1):1.
- Rainen L, Arbique JC, Asthana D, Earley MC, Geiszler RL, Krieg-Schneider F, et al. MM13-A-Collection, transport, preparation, and storage of specimens for molecular methods: approved guideline. 1nd ed. Pennsylvania,USA: CLSI; 2005. 1-51 p.
- Endler G, Slavka G, and Garland S, Tabrizi S. Stability of the specimen during preanalytics and Pathogens relevant in sexually transmitted infections. In: Kess-ler HH, editor. Molecular Diagnostics of Infectious Diseases. Berlin/New York: De Gruyter; 2010. p. 25-33, 177-84
- Youngpairoj AS, Masciotra S, Garrido C, Zahonero N, De Mendoza C, García-Lerma JG. HIV-1 drug resistance genotyping from dried blood spots stored for 1 year at 4° C. J Antimicrob Chemother. Maryam Sotoudeh Anvari et al. 17 Vol.16 No.1 Win-ter 2021 IRANIAN JOURNAL OF PATHOLOGY 2008;61(6):1217-20.
- Nechvatal JM, Ram JL, Basson MD, Namprachan P, Niec SR, Badsha KZ, et al. Fecal collection, ambient preservation, and DNA extraction for PCR amplifica-tion of bacterial and human markers from human feces. J Microbiol Methods. 2008;72(2):124-32.

Most Commented