POTENTIAL ZOONOTIC BAT-BORNE DISEASE IN INDONESIA (Part 1)
Over the past 50 years, several viruses, including Ebola virus, Marburg virus, Nipah virus, Hendra virus, severe acute respiratory syndrome coronavirus (SARS-CoV), Middle East respiratory coronavirus (MERS-CoV), and SARS-CoV-2, have been linked back to various bat species. Bats, order Chiroptera, are the only mammals capable of powered flight and are among the most ancient of mammals and underwent extensive speciation for the last 100 million years. There are currently more than 1000 species of bats, making them the second most diverse mammalian group, after rodents, and representing 20% of extant mammalian species. It is increasingly accepted that bats are important reservoirs of many known and unknown viruses, many of which could spill over into animal and human populations, including RNA viruses such as Marburg virus, Hendra virus, Sosuga virus, and Nipah virus (1,2). Bats are reported to possess efficient and varied antiviral responses associated with adaptations in their immune system and their ability to evolve. The adaptive immune mechanism in bats can suppress the pathological effects of the inflammation caused by viral infection. However, various factors, such as stress, may contribute to unbalancing the mechanism, resulting in increased viral replication and shedding and potentially becoming a source of cross-species virus transmission, including human transmission (3,4).
In addition to direct isolation of these human pathogens from bats, accumulating evidence suggests that other emerging viruses, such as Ebola viruses, severe acute respiratory syndrome coronavirus (SARS-CoV), SARS-CoV-2, and Middle East respiratory coronavirus (MERS-CoV), also originated in bats, even if other hosts, such as civets for SARS-CoV and camels for MERS-CoV, are proximate reservoirs for human infection. A growing list of emergent coronaviruses, including the Swine acute diarrhea syndrome coronavirus, which emerged from horseshoe bats and killed >20,000 pigs, and the ongoing COVID-19 pandemic, further underscores the ongoing threat of bat-borne viral emergence (1,5). The rate of emergence of novel viruses appears to be increasing due to both increased spillover from their natural reservoirs and our improved ability in detection. To date, thousands of new bat-associated viral species have been discovered from at least 28 diverse viral families, the vast majority of which are likely host specific with limited zoonotic potential. Some bat-associated viral families, such as coronaviruses, henipaviruses, lyssaviruses, and filoviruses, are diverse and of great public and veterinary health concern because of their rapid evolutionary rate, pathogenicity in human or other hosts, and proven ability to emerge. Of note, large parts of the bat virus diversity remain uncharacterized, and discovery efforts have prioritized virus families with known zoonotic potential, such as the Coronaviridae (2).


The recognition of the role of bats in viral epidemics presents the risk of bats being responsible for them and thus of considering their eradication as a solution to the risk of infection. This reaction, which is unfortunately intuitive, would prove to be totally inappropriate and even prejudicial to human health. It has already been tested in Uganda, where, as part of campaigns to prevent Marburg virus infections, the destruction of fruit bats has been carried out in some mines (6,7). This has resulted in the re-invasion of these sites by susceptible bats and multiple reintroductions of the virus into newly connected populations. Reacting like this means forgetting that Chiropterans are key species in the functioning of ecosystems. In tropical environments, they play a significant role in the pollination of plants and the long-distance dissemination of seeds. As for insectivorous bats, they play a major role in regulating insect populations and thus reduce the use of pesticides. Very sensitive to changes in their environment, bats are excellent indicators of the health of our environment. The causes of epidemics are rather to be found in the disruption of natural ecosystems inflicted by human activities: intensification of agricultural practices leading to deforestation and habitat fragmentation, habitat degradation, and rapid urbanization (7).
Bats constitute a substantial portion of mammalian diversity throughout the Asian tropics. Indonesia supports high bat diversity which is at least 200 species. It is probable that Indonesia’s abundant biodiversity of natural reservoir hosts (including bats), high tropical deforestation rates, thriving wildlife trade and hunting networks, and growing human population may increase the risk for zoonotic disease emergence (2,7–10). Therefore, this article will describe potential zoonosis spillover events in Indonesia of three viral family classes: coronaviridae, filoviridae, and paramyxoviridae.
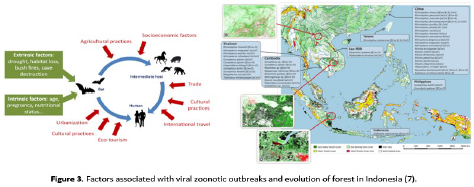

Coronavirus
Coronaviruses that circulate in bat populations have spilled over into human populations several times, and most likely will continue to be a public health threat. The diversity and broad geographical distribution of bats, the ubiquitous shedding of coronaviruses from bat populations and the molecular interactions of coronaviruses facilitate their zoonotic capacity. Coronaviruses are a diverse group of viruses infecting many different animals, and they can cause mild to severe respiratory infections in humans. Coronaviridae is subdivided into four genera, viz., Alphacorona-virus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus. Bats are important natural hosts of alphacoronaviruses and beta coronaviruses. Little is known about the specific conditions of coronavirus spillovers, but human behaviors that may increase viral exposure include activities such as bat hunting and consumption, guano farming and wildlife trading. Coronavirus shed-ding in horseshoe bats was higher in human-dominated landscapes than in natural landscapes. In addition, the legal and illegal wildlife trade results in viruses being transported over longer distances within hosts maintained in stressful and unsanitary conditions, likely increasing shedding and transmission (8,11,12).
Combined ecological and epidemiological data with modeling of the spread of bats to map the risk of exposure to bat corona-viruses across China and Southeast Asia and estimate an average of 400,000 people are infected with a SARSr-CoV every year (unreported spillover events), which and suggest that human exposure to and spillover of SARSr-CoVs may be substantially underestimated, and is undetected by surveillance programs and clinical studies in the majority of cases. Their analysis identified regions in southern China, northeastern Myanmar, Lao PDR, and northern Vietnam as having the highest diversity of SARSr-CoV bat host species. These hotspots of SARSr-CoV bat reservoir host

diversity may be particularly fruitful sites for viral discovery of novel SARSr-CoVs, assuming that viral diversity scales with host species diversity. While the data suggest significant levels of exposure, many of the diverse viral strains that infect people in the region each year may not be able to replicate well in people, cause illness, or be transmitted sufficiently among people to cause an outbreak. However, given the relatively large number of people likely to be infected each year with bat-CoVs, it is plausible that illnesses or clusters of cases due to novel bat-CoV infection occur regularly within the region, and are either not reported, or otherwise missed by clinical surveillance (10).
Among the many questions unanswered for the COVID-19 pan-demic are the animal origin and cross-species infection route of SARS- CoV-2 are yet to be uncovered. Fundamental knowledge gaps remain about the understanding of mechanisms leading to successful spillover event. To date, the closest relatives to SARS-CoV-2 is RaTG13, sampled from a Rhinolophus affinis (horseshoe bat in Yunnan province), which shared a ~96% identical overall to SARS-CoV-2. However, its spike diverges in the RBD, which suggests that it may not bind efficiently to human ACE2. In contrast, some pangolin coronaviruses exhibit strong similarity to SARS-CoV-2 in the RBD, including all six key RBD residues. Neither the bat beta coronaviruses nor the pangolin beta coronaviruses sam-pled thus far have polybasic cleavage sites. A study reported molecular and serological evidence of SARS-CoV-2 related coronaviruses (SC2r-CoVs) actively circulating in bats in Southeast Asia. Whole-genome sequences were obtained from five independent bats (Rhinolophus acuminatus) in a Thai cave yielding a single isolate (named RacCS203) which is most related to the RmYN02 isolate found in Rhinolophus malayanus in Yunnan, China (12–14).
Indonesia is reported to have 81 species of bats that belong to suborder Megachiroptera (fruit nectar-eating bats) and 158 species in the suborder Microchiroptera (non-fruit-nectar-eating bats). As many as 10 families of bats have been reported in Indonesia, including Pteropodidae, which belongs to the Meg-achiroptera suborder, and Rhinopomatidae, Emballonuridae, Nycteridae, Megadermatidae, Rhinolophidae, Hiposideridae, Vespertilionidae, Minioptereridae, and Molossidae families that belong to the Microchiroptera suborder. As an archipelago country, Indonesia has the highest number of fruit- and nectar-eating bat species in the world. Fruit-nectar bats are very important for the process of pollination, fertilization, and seed dispersal. There are at least 21 genera and 81 species that belong to Pteropodi-dae family. The discovery of BatCoV in several species of bats in Indonesia suggests that bats are potential natural reservoir hosts for coronavirus and may transmit that virus to other species, such as occurred in previous infectious disease outbreaks such as SARS-CoV and MERS-CoV. Recent study in Indonesia collected a total of 182 rectal swab samples from 4 species of bats were acquired from bat collectors who were going to sell the bats to restaurants for consumption and from animal markets in several regencies/cities in Central Java Province (Surakarta City, Mage-lang Regency), Yogyakarta Province, and West Java Province (Bogor City, Cianjur Regency) in 2020 and 2021. They identified the presence of BatCoV on Cynopterus brachyotis, Macroglossus minimus, and Rousettus amplexicaudatus. The results showed that the BatCoV included in this study are from an unclassified coronavirus group. Notably, SARS-CoV-2 viral RNA and antibodies were not detected in the sampled bats (9). However, close contacts between humans, bats, and other animals have a high potential for transmitting zoonotic diseases. Further studies in Indonesia regarding coronaviruses carried by bats or other animals and the possible effects of environmental conditions are needed to identify possible novel virus transmission routes, particularly in live animal markets. Early detection of pathogen transmission and the application of appropriate control measures may minimize the destructive impacts on global health. Thus, more surveillance studies are needed to investigate the potentially important role of bats as natural reservoir hosts in the inter-species transmission of coronaviruses.
-Continued to part 2 in February 2022-
References:
- Wang L-F, Anderson DE. Viruses in bats and potential spillo-ver to animals and humans. Curr Opin Virol. 2019 Feb;34:79–89.
- Letko M, Seifert SN, Olival KJ, Plowright RK, Munster VJ. Bat-borne virus diversity, spillover and emergence. Nat Rev Mi-crobiol [Internet]. 2020;18(8):461–71. Available from: https://doi.org/10.1038/s41579-020-0394-z
- Subudhi S, Rapin N, Misra V. Immune System Modulation and Viral Persistence in Bats: Understanding Viral Spillover. Viruses [Internet]. 2019 Feb 23;11(2):192. Available from: https://pubmed.ncbi.nlm.nih.gov/30813403
- Banerjee A, Baker ML, Kulcsar K, Misra V, Plowright R, Moss-man K. Novel Insights Into Immune Systems of Bats [Internet]. Vol. 11, Frontiers in Immunology . 2020. p. 26. Available from: https://www.frontiersin.org/article/10.3389/fimmu.2020.00026
- Allocati N, Petrucci AG, Di Giovanni P, Masulli M, Di Ilio C, De Laurenzi V. Bat-man disease transmission: zoonotic patho-gens from wildlife reservoirs to human populations. Cell death Discov. 2016;2:16048.
- Gibb R, Redding DW, Chin KQ, Donnelly CA, Blackburn TM, Newbold T, et al. Zoonotic host diversity increases in human-dominated ecosystems. Nature [Internet]. 2020;584(7821):398–402. Available from: https://doi.org/10.1038/s41586-020-2562-8
- Afelt A, Frutos R, Devaux C. Bats, Coronaviruses, and Defor-estation: Toward the Emergence of Novel Infectious Diseas-es? [Internet]. Vol. 9, Frontiers in Microbiology . 2018. p. 702. Available from: https://www.frontiersin.org/article/10.3389/fmicb.2018.00702
- Ruiz-Aravena M, McKee C, Gamble A, Lunn T, Morris A, Snedden CE, et al. Ecology, evolution and spillover of coro-naviruses from bats. Nat Rev Microbiol [Internet]. 2021; Available from: https://doi.org/10.1038/s41579-021-00652-2
- Dharmayanti NLPI, Nurjanah D, Nuradji H, Maryanto I, Ex-ploitasia I, Indriani R. Molecular detection of bat corona-viruses in three bat species in Indonesia. J Vet Sci [Internet]. 2021 Nov;22(6):0. Available from: https://doi.org/10.4142/jvs.2021.22.e70
- Sánchez CA, Li H, Phelps KL, Zambrana-Torrelio C, Wang L-F, Olival KJ, et al. A strategy to assess spillover risk of bat SARS related coronaviruses in Southeast Asia. medRxiv : the preprint server for health sciences. 2021
- Zhou Z, Qiu Y, Ge X. The taxonomy, host range and pathogenicity of coronaviruses and other viruses in the Nidovi-rales order. Anim Dis [Internet]. 2021;1(1):5. Available from: https://doi.org/10.1186/s44149-021-00005-9
- Kuldeep D, Sharun K, Ruchi T, Shubhankar S, Sudipta B, Singh MY, et al. Coronavirus Disease 2019–COVID-19. Clin Microbiol Rev [Internet]. 2022 Jan 11;33(4):e00028-20. Avail-able from: https://doi.org/10.1128/CMR.00028-20
- Delaune D, Hul V, Karlsson EA, Hassanin A, Ou TP, Baidaliuk A, et al. A novel SARS-CoV-2 related coronavirus in bats from Cambodia. Nat Commun [Internet]. 2021;12(1):6563. Available from: https://doi.org/10.1038/s41467-021-26809-4
- Wacharapluesadee S, Tan CW, Maneeorn P, Duengkae P, Zhu F, Joyjinda Y, et al. Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia. Nat Commun [Internet]. 2021;12(1):972. Available from: https://doi.org/10.1038/s41467-021-21240-1

Most Commented